Random walk vectors for clustering (part II – perspective switch)
This post is a second part of the series of posts that will result in combining random walk vectors for clustering. So far we have understood what the similarity is, how to build similarity matrix based on distance matrix. We know the similarity matrix is a structure that encodes similarities between all objects in our dataset.
Today we will further motivate our quest for similarity matrix started in the previous post. We will tell a little bit about what the graph is and how to switch from point cloud perspective to graph perspective. The bridge between these two worlds is the similarity matrix introduced last time.
Basics first.
A wide range of structures occurring in nature are naturally expressed as networks. Whenever a set of objects and relations among them appears, the whole setting can be interpreted this way. A typical example of such network is – becoming hot topic in last decade – social network. A node in that type of a network is an abstraction of a social unit (mainly a person) and an edge represents social relation between pairs of nodes like friendship, business relation, co-interest etc. Another type of network is the internet itself where websites are represented as nodes and links between websites as connections between them. In fact, any set of objects with relation between them can be modeled as a network.
Formally a network is called graph and it consists of set of nodes and set of edges and is usually formalized as a pair \(G = (V,E)\) where \(V\) is a set of \(N\) objects indexed with integers \({1, 2, \dots, N }\) and \(E\) is a function:
\[E : V^2 \rightarrow \mathcal{R}\]In practice it means that for each pair of nodes we have a number that expresses strength of underlying relation. Depending on properties of \(E\) graph can be considered
- undirected – if \( E(a,b) = E(b,a) \) for all nodes, meaning underlying relation is modeled as symmetric (like friendship on Facebook)
- directed – if \( E(a,b) = E(b,a) \) does not hold for at least one node pair, meaning underlying relation is modeled as not symmetric (follower on Twitter)
- unweighted – if range of E values is \({0,1}\), meaning underlying relation is binary (same family membership, again)
- weighted – if range of E values exceeds \({0,1}\), meaning underlying relation strength may have various values (family relation strength)
One convenient way of describing a graph is by using a matrix. Different names are used for the matrix under consideration, here we will call it “Weight matrix” \(\mathrm{W}\). The matrix encodes function \(E\) values in the following way:
\[E(a,b) = \mathrm{W}_{ij}\]where \(i\) and \(j\) are indices of objects \(a\) and \(b\) respectively. And, once again, depending on graph nature (whether it is directed, undirected, unweighted or weighted) underlying weight matrix reflects it. It is symmetric if graph is undirected or binary in case of unweighted graph etc.
One important note here. Matrix \(\mathrm{W}\) and a graph \(G\) can be used interchangeably. In other words matrix \(\mathrm{W}\) encodes all information about given graph \(G\). And that is the important piece of information for us, in the context we are now.
Why is this important bit? Please recall what we did last time, we encoded given dataset in a form of similarity matrix. If we now treat similarity as a relation, our objects as nodes and similarity matrix as matrix \(\mathrm{W}\) we can encode our dataset as a graph. We can change the point of view and think of our framework as if we were given a graph.
Why do we need such a perspective switch? We gain much by doing so. Graph theory is a very popular field of research therefore we gain the theories and tools from the world of graphs that we can later apply to our original problem.
Random walk on a graph – its interpretation as similarity
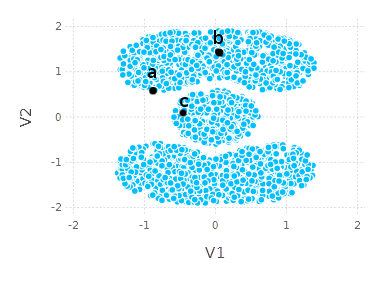
Let us now consider a graph \(G\) given by data similarity matrix \(W\) and answer few simple questions. Given a single data point (indexed with \(i\)), how would you find \(k\) most similar objects? The very first idea is to look at \(i\)-th column of \(W\) matrix, find \(k\) highest values (so in fact just to pick \(k\) most similar objects according to similarity measure), and we are done. It is fine, it is a good solution, but let us take a look at the example below:
Would you consider \(a\) similar to \(c\) more than \(a\) to \(b\)? It is not that obvious. If I were asked I would say that \(a\) and \(b\) belong together more than \(a\) and \(c\). If we consider points’ neighborhood and global shape of dataset \(a\) might be considered closer to \(b\) than to \(c\).
What can we do to express these kinds of similarities? One idea is to spread similarity information across the graph. You can imagine a system of nodes and pipes and water flow between nodes through the pipes. We start the water flow from the single node. The amount of water that goes through any pipe is proportional to similarity (between nodes). If water flow like that lasts for “some time” we get similarity on a “more global” scale. The amount of water in nodes may then express similarity.
If you consider that scenario (if similarity matrix is parameterized properly) and apply it to our example above, you can imagine that starting from node \(a\) after few steps of such a flow the amount of water in \(b\) would be greater than in \(c\).
Formally we describe presented “water flow” as a random walk on graph. Water here is replaced by probability and system of pipes is replaced by transition matrix. Time is discrete in such a setting.
Let us define transition matrix \(\mathrm{P}\) based on given similarity matrix \(W\).
\[\mathrm{P} = \mathrm{W} \mathrm{D}^{-1}\]where \(\mathrm{D} = \mathrm{diag}(\mathrm{W} \cdot \vec{1})\) (diag is a diagonal matrix constructor).
\(\mathrm{P}\) is a column stochastic matrix – it means values in each column sum up to 1. To model the flow over the network we need a notion of seed distribution. Seed distribution is just an initial “water” distribution in our pipes system. Formally it is given by vector \(v_0\) restricted to
\[\sum_{i}^{D} v_{0i} = 1\]Having \(\mathrm{P}\) and \(v_{0}\) let’s define a random walk vector at time \(k\) as
\[v_{k} = \mathrm{P}^k v_{0}\]It reflects how our pipes system would look like (in terms of water amount in nodes) after \(k\) steps (or after \(k\) time units passed).
Let us get back to our abstraction – to get similarities that reflect global “shape” of our dataset – we can just perform some number of steps of a random walk and treat resulting random walk vector as a similarity vector. That hopefully solves the problem of similarities between \(a, b\) and \(c\) in our toy example.
Random walk in Julia
Let’s try to implement random walk and check if that is going to help us in case of cassini dataset to model similarities based on dataset ‘shape’.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
using Gadfly
using DataFrames
using Distances
cassini_data_frame = readtable("cassini.csv")
cassini_data_matrix = transpose(convert(Array{Float64,2}, cassini_data_frame[:,2:3]))
function gaussian_similarity(sigma)
return x -> exp((-x.^2)./(2*sigma^2))
end
distance_matrix = pairwise(Euclidean(), cassini_data_matrix)
similarity_matrix = gaussian_similarity(0.2)(distance_matrix)
P = (similarity_matrix) * diagm(1. ./ (similarity_matrix * ones(size(similarity_matrix,2))))
plot(x = cassini_data_matrix[1,:], color=(P*v), y = cassini_data_matrix[2,:], Geom.point)
plot(x = cassini_data_matrix[1,:], color=(P^2*v), y = cassini_data_matrix[2,:], Geom.point)
plot(x = cassini_data_matrix[1,:], color=(P^4*v), y = cassini_data_matrix[2,:], Geom.point)
plot(x = cassini_data_matrix[1,:], color=(P^8*v), y = cassini_data_matrix[2,:], Geom.point)
plot(x = cassini_data_matrix[1,:], color=(P^16*v), y = cassini_data_matrix[2,:], Geom.point)
plot(x = cassini_data_matrix[1,:], color=(P^32*v), y = cassini_data_matrix[2,:], Geom.point)
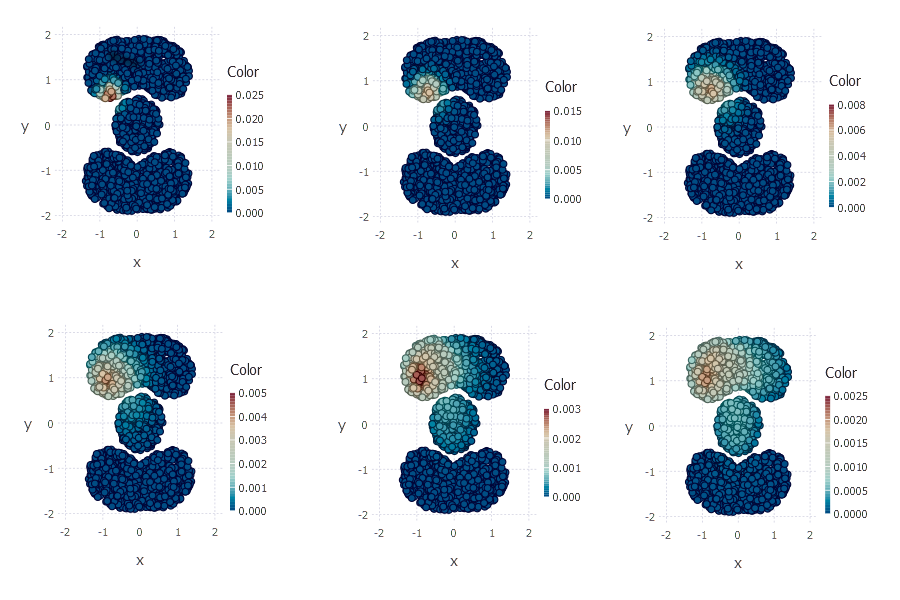
On the right bottom plot you can see random walk vector after 32 steps. As you can see random walk vector captures shape of dataset such that point \(a\) is closer to \(b\) than to \(c\).
How can we now interpret random walk vector? One way is to think of it as a vector reflecting similarities between points such that points with high corresponding probabilities should be considered close to each other. Each vector expresses points’ participation in local shape being discovered around given starting point. The trick I would like to use here is to consider many random walk vectors (with different starting nodes – seed vectors) at a time. Each vector would then express points’ participation in local shapes being discovered. The main idea is to treat such participation as a new dimension and represent points in a new space that is defined by points’ participations in different regions of a given dataset. Of course few questions may be asked directly, how to choose starting points, how many steps should be taken, how to choose sigma parameter, etc.
In the next post we will try to represent one example dataset in a space briefly defined above. We will also give some basic algebraic explanation of what we will try to do. We will also try to write some more Julia code!