Large Scale Spectral Clustering with Landmark-Based Representation (in Julia)
In this post we will implement and play with a clustering algorithm of a mysterious name Large Scale Spectral Clustering with Landmark-Based Representation (or shortly LSC – corresponding paper here). We will first explain the algorithm step by step and then map it to Julia code (github link).
Spectral Clustering
Spectral clustering (wikipedia entry) is a term that refers to many different clustering techniques. The core of the algorithm does not differ though. In essence, it is a method that relies on spectrum (eigendecomposition) of input data similarity matrix (or its transformations). Given input dataset encoded in a matrix \(X\) (such that each single data entry is a column of that matrix) – spectral clustering requires a similarity (adjacency in case of graphical interpretation) matrix \(A\) where
\[A_{ij} = f(X_{\cdot i}, X_{\cdot j})\]and function \(f\) is some measure of similarity between data points.
One specific spectral clustering algorithm (Ng, Jordan, and Weiss 2001) relies on matrix \(W\) given by
\[W = D^{-1/2} A D^{-1/2}\]where \(D\) is a diagonal matrix with diagonal elements \((i,i)\) corresponding to sum of \(i\)-th row of \(A\).
In Jordan, Weiss algorithm first \(k\) (with largest eigenvalue) eigenvectors of \(W\) are computed and (after normalization) they go as input to k-means to produce final dataset partition (clustering). Same idea motivates authors of LSC. The core idea of the LSC method is to limit the complexity of EVD of \(W\).
Large Scale Spectral Clustering with Landmark-Based Representation
One possible way to go to limit EVD complexity is to reduce the original problem to another (easier) one. In LSC paper authors construct matrix similar to \(W = D^{-1/2} A D^{-1/2}\) in a clever way. Matrix \(W\) used in LSC has the following form:
\[W = \hat{Z}^{T} \hat{Z}\](authors construct \(\hat{Z}\) such that it is sparse matrix of dimensionality much lower than original matrix)
Therefore it is sufficient enough to compute Singular Value Decomposition of sparse small matrix \(\hat{Z}\) instead of computing EVD of \(W\). Please recall, SVD of any matrix \(X\) produces eigenvectors of both \(XX^T\) and \(X^TX\)
Construction of matrix \(\hat{Z}\)
Matrix \(\hat{Z}\) is defined as:
\[\hat{Z} = D^{-1/2} Z\]where \(Z\) is a sparse matrix that can be seen as new input data \(X\) representation and again \(D^{-1/2}\) is diagonal matrix with row sums entries of \(Z\). Columns of matrix \(Z\) are defined as coefficients of a weighted sum (linear combination) of a centroid vectors given by prior k-means clustering (or randomly chosen input vectors!). These vectors are called Landmarks
Here is exact routine to get matrix \(Z\) (assuming choice based on k-means) :
- Given input data matrix \(X\) compute \(p\) k-means centroids of \(X\) and store them in \(U\)
- For each data point \(x_i\) produce sparse vector \(z_{i}\) with non zero entries corresponding to the closest \(r\) clustering centroids from \(U\) given by gaussian kernel
- Normalize \(z_{i}\) and store in matrix \(Z\) as an \(i\)-th column
Computing eigenvectors of matrix \(W\) – spectral clustering core part
Having \(W\) constructed according to aforementioned formulas, the next core part of the algorithm is to compute EVD of \(W\). As was mentioned, eigendecomposition is not computed directly on \(W\) as this would be inefficient. Instead of direct EVD on \(W\) SVD of sparse and smaller \(\hat{Z}\) is computed. SVD of \(\hat{Z}\) is given by:
\[\hat{Z} = A \Sigma B^T\](note that it is of course different \(A\) than similarity matrix above)
Here authors compute matrix \(A\) from SVD fast to finally compute \(B\) according to the formula:
\[B^T = \Sigma^{-1} A^T \hat{Z}\]Matrix \(B\) contains our final eigenvectors that are input for final k-means clustering. (our implementation skips that as svd routine in Julia computes \(B\) directly).
Final step: k-means on eigenvectors
Now having eigenvectors of \(W\) authors apply classical k-means algorithm to get final data clustering. The main gain here is the computational complexity reduction. Normally, eigendecomposition has complexity of \(O(n^3)\) in data set cardinality \(n\). After a bit of gymnastics as above they achieve \(O(p^3 + p^2n)\) where \(p\) is number of centroids from initial k-means (or number of randomly selected landmarks in case of such algorithm variant)
LSC – implementation
Let’s start with routine to get \(p\) landmarks.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
using Clustering;
using Distances;
function getLandmarks(X, p, method=:Random)
if(method == :Random)
numberOfPoints = size(X,2);
landmarks = X[:,randperm(numberOfPoints)[1:p]];
return landmarks;
end
if(method == :Kmeans)
kmeansResult = kmeans(X,p)
return kmeansResult.centers;
end
throw(ArgumentError('method can only be :Kmeans or :Random'));
end
We also need a function to compute \(\hat{Z}\)
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
function gaussianKernel(distance, bandwidth)
exp(-distance / (2*bandwidth^2));
end
function composeSparseZHatMatrix(X, landmarks, bandwidth, r)
distances = pairwise(Distances.Euclidean(), landmarks, X);
similarities = map(x -> gaussianKernel(x, bandwidth), distances);
ZHat = zeros(size(similarities));
for i in 1:(size(similarities,2))
topLandMarksIndices = selectperm(similarities[:,i], 1:r, rev=true);
topLandMarksCoefficients = similarities[topLandMarksIndices, i];
topLandMarksCoefficients = topLandMarksCoefficients / sum(topLandMarksCoefficients);
ZHat[topLandMarksIndices,i] = topLandMarksCoefficients;
end
return diagm(sum(ZHat,2)[:])^(-1/2) * ZHat;
end
Finally, we can implement core LSC routine as
1
2
3
4
5
6
7
function LSCClustering(X, nrOfClusters, nrOfLandmarks, method, nonZeroLandmarkWeights, bandwidth)
landmarks = getLandmarks(X, nrOfLandmarks, method)
ZHat = composeSparseZHatMatrix(X, landmarks, bandwidth, nonZeroLandmarkWeights)
svdResult = svd(transpose(ZHat))
clusteringResult = kmeans(transpose(svdResult[1])[:1:nrOfClusters],nrOfClusters);
return clusteringResult
end
Please note that our implementation differs a bit from the original one (here, we consider first \(k\) eigenvectors as new data representation).
LSC - toy data examples
Here is the result of LSC clustering (with k-means landmark generation) on toy datasets - they all have cardinality 2000
(Datasets are available to download: cassini, shapes, smiley and spirals)
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
using Gadfly
# smiley example
smiley = readtable("smiley.csv");
smiley = transpose(convert(Array{Float64,2}, smiley[:,1:2]))
result = LSCClustering(smiley, 4, 50, :Kmeans, 4, 0.4)
plot(x = smiley[1,:], y = smiley[2,:], color = result.assignments)
# spirals example
spirals = readtable("spirals.csv");
spirals = transpose(convert(Array{Float64,2}, spirals[:,1:2]))
result = LSCClustering(spirals, 2, 150, :Kmeans, 4, 0.04)
plot(x = spirals[1,:], y = spirals[2,:], color = result.assignments)
# shapes example
shapes = readtable("shapes.csv");
shapes = transpose(convert(Array{Float64,2}, shapes[:,1:2]))
result = LSCClustering(shapes, 4, 50, :Kmeans, 4, 0.04)
plot(x = shapes[1,:], y = shapes[2,:], color = result.assignments)
# cassini example
cassini = readtable("cassini.csv");
cassini = transpose(convert(Array{Float64,2}, cassini[:,2:3]))
result = LSCClustering(cassini, 3, 50, :Kmeans, 4, 0.4)
plot(x = cassini[1,:], y = cassini[2,:], color = result.assignments)
Obviously, even though results look quite promising we cannot pass over one problem: parameter choice. Here, we have quite a few of them - number of landmarks, \(Z\) density (via \(r\)), and Gaussian bandwidth - and sometimes (this time in particular!) choosing working set of parameters is not an easy task.
LSC - MNIST dataset
Let’s finally try LSC on a ‘more serious’ dataset - handwritten digits dataset called MNIST. In this experiment we are going to compare LSC with classical k-means. We are going to use NMI (normalized mutual information) clustering quality as our metric.
Let’s quickly implement NMI (according to this)
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
# Pkg.add("Iterators")
using Iterators
function entropy(partitionSet, N)
return mapreduce(x -> -(length(x) / N) * log(length(x) / N), +, partitionSet)
end
function getPartitionSet(vector, nrOfClusters)
d = Dict(zip(1:nrOfClusters,[Int[] for _ in 1:nrOfClusters]));
for i in 1:length(vector)
push!(d[vector[i]], i)
end
return collect(values(d))
end
function mutualInformation(partitionSetA, partitionSetB, N)
anonFunc = (x) -> (
intersection = size(intersect(partitionSetA[x[1]], partitionSetB[x[2]]),1);
return intersection > 0 ?
(intersection/N) * log((intersection * N) / (size(partitionSetA[x[1]],1) * size(partitionSetB[x[2]],1))) : 0;
)
mapreduce(anonFunc, + , product(1:length(partitionSetA),1:length(partitionSetB)))
end
function normalizedMutualInformation(partitionSetA, partitionSetB, N)
mutualInformation(partitionSetA, partitionSetB, N) / ((entropy(partitionSetA, N) + entropy(partitionSetB, N)) / 2)
end
And now we are ready to go. Let’s compare our results with k-means for a sanity check (running 50 experiments):
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
# Pkg.add("MNIST")
using MNIST;
# reading dataset (60k objects)
data, labels = MNIST.traindata(); # using testdata() instead will return smaller (10k objects) dataset
# normalizing it
data = (data .- mean(data,2)) / std(data .- mean(data,2));
nrOfExperiments = 50;
kmeansResults = [];
LSCResults = [];
for i in 1:nrOfExperiments
LSCMnistResult = LSCClustering(data, 10, 350, :Kmeans, 5, 0.5);
kmeansMnistResult = kmeans(data,10);
kmeansNMI = normalizedMutualInformation(getPartitionSet(kmeansMnistResult.assignments, 10) , getPartitionSet(labels + 1 ,10), 60000)
LSCNMI = normalizedMutualInformation(getPartitionSet(LSCMnistResult.assignments, 10) , getPartitionSet(labels + 1 ,10), 60000)
push!(kmeansResults, kmeansNMI);
push!(LSCResults, LSCNMI);
end
# Pkg.add("DataFrames");
# Pkg.add("Gadfly");
using DataFrames;
using Gadfly;
df = DataFrame(nmi = [kmeansResults; LSCResults], algo = [["kmeans" for _ in 1:50];["LSC" for _ in 1:50]]);
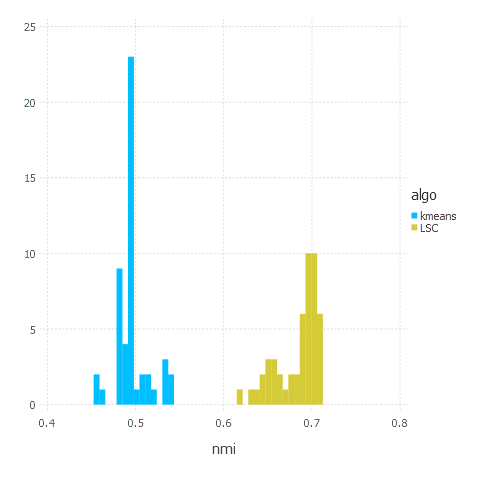
plot(df, x="nmi", color="algo", Geom.histogram(bincount=40));
Summary
That’s it. LSC clustering algorithm turned out to be better in terms of quality (as promised in a paper) than k-means. The main challenge though was proper parameters’ values choice - usually we had to tweak Gaussian kernel bandwidth and number of landmarks a bit to make the algorithm produce good results. K-means does not suffer from that problem so it is usually ‘practically’ faster - as you don’t need to experiment with parameters (which is a hard problem itself) to get good clustering.